Anisotropic data with scale#
Display a 3D image with anisotropic voxel spacing using the scale
parameter so that the volume appears with correct proportions.
Microscopy data is often anisotropic: the voxel spacing differs across
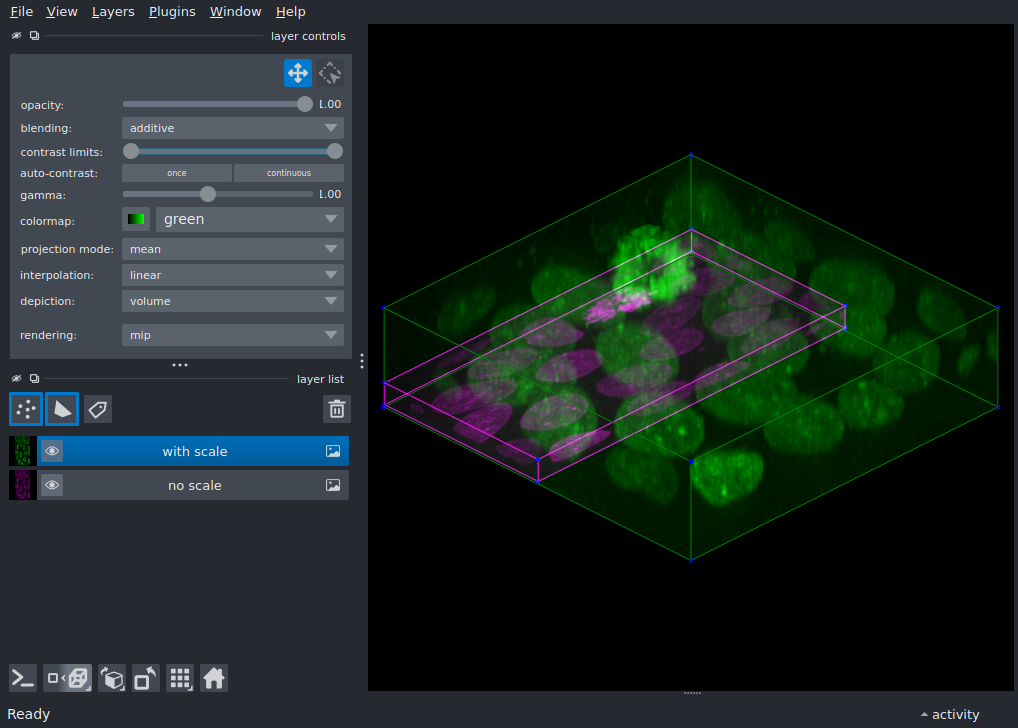
axes. Without scale, napari treats every voxel as a unit cube and
the volume appears with incorrect proportions.
Setting scale to the real voxel dimensions corrects this.
Toggle the visibility of each layer to compare the difference.
from skimage import data
import napari
# cells3d has voxel spacing approximately (0.29, 0.26, 0.26) in (z, y, x).
# We subsample z by 4 and x by 2 to simulate more strongly anisotropic data.
# After subsampling, the effective voxel spacing becomes (1.16, 0.26, 0.52).
# The ratio is approximately (4.5, 1, 2), which we pass to ``scale``.
cells = data.cells3d()
nuclei = cells[::4, 1, :, ::2]
scale = (4.5, 1, 2)
viewer = napari.Viewer(ndisplay=3)
viewer.add_image(
nuclei,
name='no scale',
blending='additive',
colormap='magenta',
)
viewer.add_image(
nuclei,
name='with scale',
blending='additive',
colormap='green',
scale=scale,
)
viewer.layers['no scale'].bounding_box.line_color = 'magenta'
viewer.layers['no scale'].bounding_box.visible = True
viewer.layers['with scale'].bounding_box.line_color = 'green'
viewer.layers['with scale'].bounding_box.visible = True
viewer.camera.angles = (-45, 0, -60)
viewer.fit_to_view()
if __name__ == '__main__':
napari.run()